set.seed(577)
n <- 200; p <- 50
X <- matrix(rnorm(n * p), n, p)
beta_true <- c(rep(2, 5), rep(0, 45))
y <- X %*% beta_true + rnorm(n, sd = 2)
dat <- data.frame(y = y, X)
# Manual 10-fold CV: compare models using first d predictors (d = 1,...,15)
k <- 10
folds <- sample(rep(1:k, length.out = n))
n_vars <- 1:15
cv_errors <- matrix(NA, k, length(n_vars))
for (d in seq_along(n_vars)) {
pred_names <- paste0("X", 1:n_vars[d])
fmla <- as.formula(paste("y ~", paste(pred_names, collapse = " + ")))
for (j in 1:k) {
train_idx <- which(folds != j)
test_idx <- which(folds == j)
fit <- lm(fmla, data = dat[train_idx, ])
pred <- predict(fit, newdata = dat[test_idx, ])
cv_errors[j, d] <- mean((dat$y[test_idx] - pred)^2)
}
}
cv_mean <- colMeans(cv_errors)
cv_se <- apply(cv_errors, 2, sd) / sqrt(k)
cv_results <- tibble(
n_predictors = n_vars,
cv_mean = cv_mean,
cv_lo = cv_mean - cv_se,
cv_hi = cv_mean + cv_se
)
p_kfold <- ggplot(cv_results, aes(x = n_predictors, y = cv_mean)) +
geom_ribbon(aes(ymin = cv_lo, ymax = cv_hi), alpha = 0.2, fill = colors["blue"]) +
geom_line(linewidth = 1.2, color = colors["blue"]) +
geom_point(size = 3, color = colors["blue"]) +
geom_vline(xintercept = n_vars[which.min(cv_mean)], linetype = "dashed", color = colors["red"]) +
annotate("text", x = n_vars[which.min(cv_mean)] + 0.3, y = max(cv_mean) * 0.97,
label = paste0("Optimal: ", n_vars[which.min(cv_mean)], " predictors"),
color = colors["red"], hjust = 0, size = 4.5) +
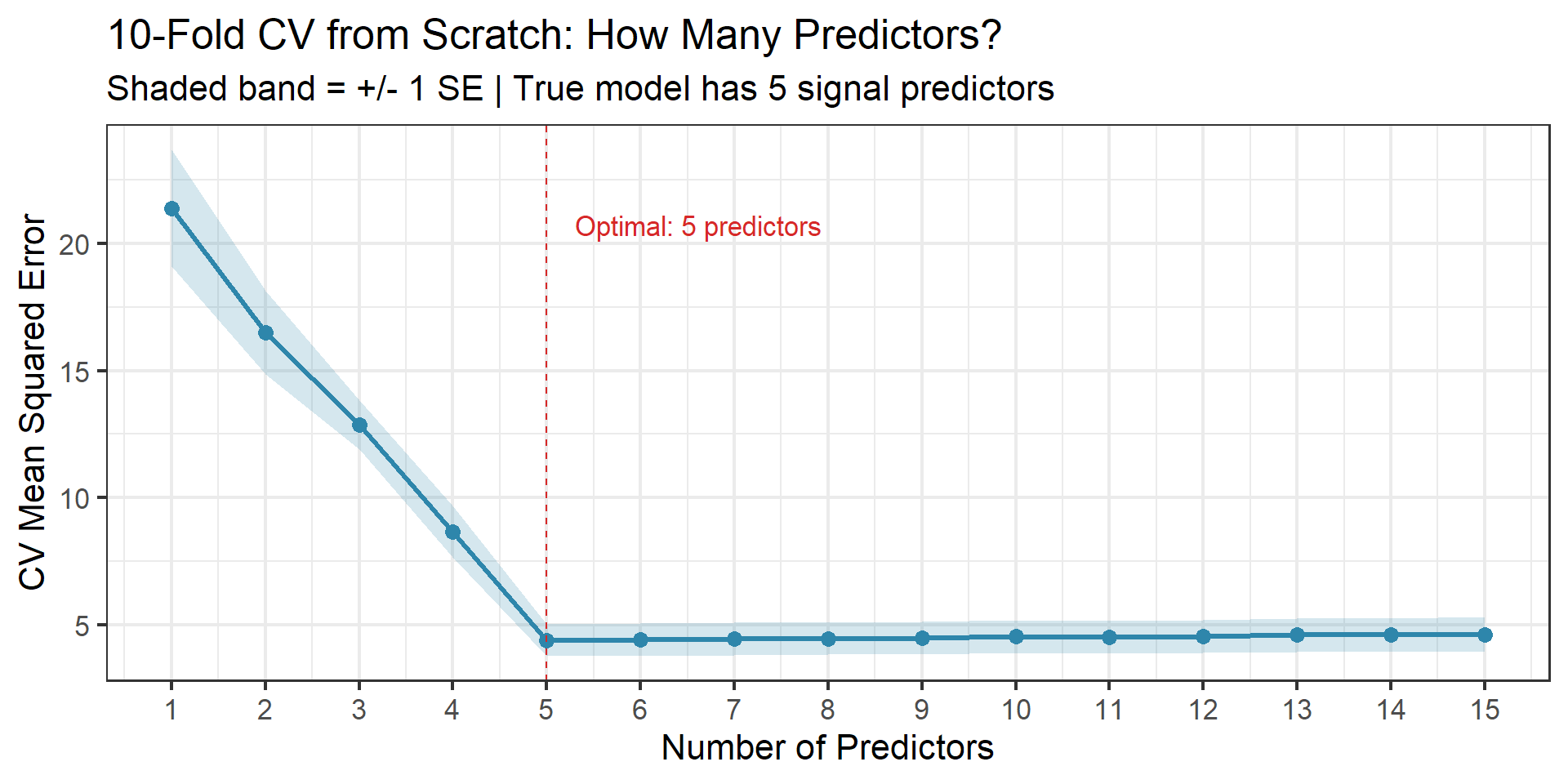
labs(title = "10-Fold CV from Scratch: How Many Predictors?",
subtitle = "Shaded band = +/- 1 SE | True model has 5 signal predictors",
x = "Number of Predictors", y = "CV Mean Squared Error") +
scale_x_continuous(breaks = n_vars) +
theme_bw(base_size = 16)
ggsave(file.path(fig_dir, "07_kfold_scratch.png"), p_kfold, width = 10, height = 5, dpi = 150)
p_kfoldWeeks 4–5: Resampling, Model Selection & Conformal Prediction
Stat 577 — Statistical Learning Theory
Spring 2026
The Information-Theoretic Core
The Problem We Have Been Ignoring
Last week, we kept saying “choose \(\lambda\) or other such tuning parameters, by cross-validation.”
We had 5+ tuning parameters and no tools to pick them.
Today we fix that.
We will build a chain of results connecting:
- Information theory (AIC)
- Bayesian model evidence (BIC)
- Data splitting (cross-validation)
- Resampling (bootstrap)
- Distribution-free prediction (conformal inference)
The Punchline (Spoiler)
These seemingly unrelated philosophies often give the same answer. Understanding why is one of the most satisfying parts of statistical learning theory.
Why Training Error Lies
Recall the bias-variance picture from Week 1:
- Training error always decreases with model complexity
- Test error is U-shaped
Training error is optimistic — it evaluates \(\hat{f}\) on the same data used to build it.
Define:
- \(\overline{\text{err}}\) = training error (average loss over training set)
- \(\text{Err}_{\text{in}}\) = in-sample error (expected loss at training \(X\) values, new \(Y\))
The optimism of training error:
\[\text{op} = \text{Err}_{\text{in}} - \overline{\text{err}}\]
Quantifying the Optimism
The Covariance Penalty (ESL Theorem 7.4)
\[E[\text{op}] = \frac{2}{n}\sum_{i=1}^{n} \text{Cov}(\hat{y}_i, y_i)\]
For a linear smoother \(\hat{\mathbf{y}} = \mathbf{S}\mathbf{y}\):
\[E[\text{op}] = \frac{2d \cdot \sigma^2}{n}, \qquad d = \text{tr}(\mathbf{S})\]
Key steps of the derivation:
- Write \(\text{Err}_{\text{in}} - \overline{\text{err}}\) in terms of expectations
- The cross-term telescopes into covariances
- For linear smoothers: \(\text{Cov}(\hat{y}_i, y_i) = S_{ii}\sigma^2\), so sum gives \(\text{tr}(\mathbf{S})\sigma^2\)
Interpretation: Complex models (large \(d\)) have more optimism — their training error is a worse guide to true performance.
Information Theory and Statistical Learning
Why Information Theory?
At first glance, information theory and statistics seem like different fields. One is about compressing signals; the other is about learning from data.
But they are deeply intertwined — and understanding why is essential for modern statistical learning.
The core question in both fields is the same:
How much do we learn from observing data?
- Shannon (1948): How many bits do we need to encode a message?
- Fisher (1925): How much information does a sample carry about a parameter?
- Kullback & Leibler (1951): How do we measure the “distance” between what we believe and what is true?
These are not analogies — they are the same mathematics viewed from different angles.
Information: From Bits to Inference
Shannon’s key insight: The information content of an event with probability \(p\) is \(\log(1/p)\).
- A coin flip (\(p = 1/2\)): \(\log_2(2) = 1\) bit of information
- A rare event (\(p = 1/1000\)): \(\log_2(1000) \approx 10\) bits — rare events are more “surprising”
- A certain event (\(p = 1\)): \(\log_2(1) = 0\) bits — no surprise, no information
Entropy: Average Information Content
\[H(p) = -\sum_y p(y) \log p(y) = E_p\!\left[\log \frac{1}{p(Y)}\right]\]
Entropy measures the average surprise from drawing from \(p\). It is maximized when \(p\) is uniform (maximum uncertainty) and minimized when \(p\) is degenerate (no uncertainty).
Why statisticians care: Entropy quantifies how much uncertainty remains in a random variable. Reducing entropy by conditioning on data is, in a precise sense, learning.
From Entropy to Cross-Entropy
Suppose the truth is \(p\) but we use model \(q\) to encode messages.
The cross-entropy measures the average cost of encoding under the wrong model:
\[H(p, q) = -\sum_y p(y) \log q(y) = E_p\!\left[\log \frac{1}{q(Y)}\right]\]
Note that \(H(p, q) \geq H(p)\) always. Using the wrong model never helps.
The gap between cross-entropy and entropy is exactly the KL divergence:
\[D_{\text{KL}}(p \,\|\, q) = H(p, q) - H(p)\]
This decomposition reveals what KL divergence really measures: the excess cost of using the wrong model.
Why This Matters for ML
When we train a classifier by minimizing cross-entropy loss, we are directly minimizing KL divergence to the true data distribution (since \(H(p)\) is a constant). This is not a coincidence — it is the theoretical foundation for the most common loss function in deep learning.
How Do We Measure How Wrong a Model Is?
Before we can derive AIC, we need a way to measure the “distance” between two probability distributions.
Suppose the truth is \(p(y)\) and our model says \(q(y)\).
Intuition from coding theory:
- If you design an optimal code for distribution \(p\), messages cost \(\log(1/p(y))\) bits on average
- If you mistakenly design for \(q\), messages cost \(\log(1/q(y))\) bits on average
- The extra cost of using the wrong code is the KL divergence
KL Divergence: Definition
Kullback-Leibler Divergence
\[D_{\text{KL}}(p \,\|\, q) = E_p\left[\log \frac{p(Y)}{q(Y)}\right] = \int p(y) \log\frac{p(y)}{q(y)}\, dy\]
Properties:
- \(D_{\text{KL}}(p \,\|\, q) \geq 0\) (Gibbs’ inequality)
- \(D_{\text{KL}}(p \,\|\, q) = 0\) if and only if \(p = q\) (a.e.)
- NOT symmetric: \(D_{\text{KL}}(p \,\|\, q) \neq D_{\text{KL}}(q \,\|\, p)\) in general
- Not defined when \(q(y) = 0\) and \(p(y) > 0\) (the support of \(q\) must contain the support of \(p\))
Not a True Distance
KL divergence fails the triangle inequality and symmetry. It is a divergence, not a metric. Think of it as a directed measure of surprise.
KL Divergence: Visual Intuition
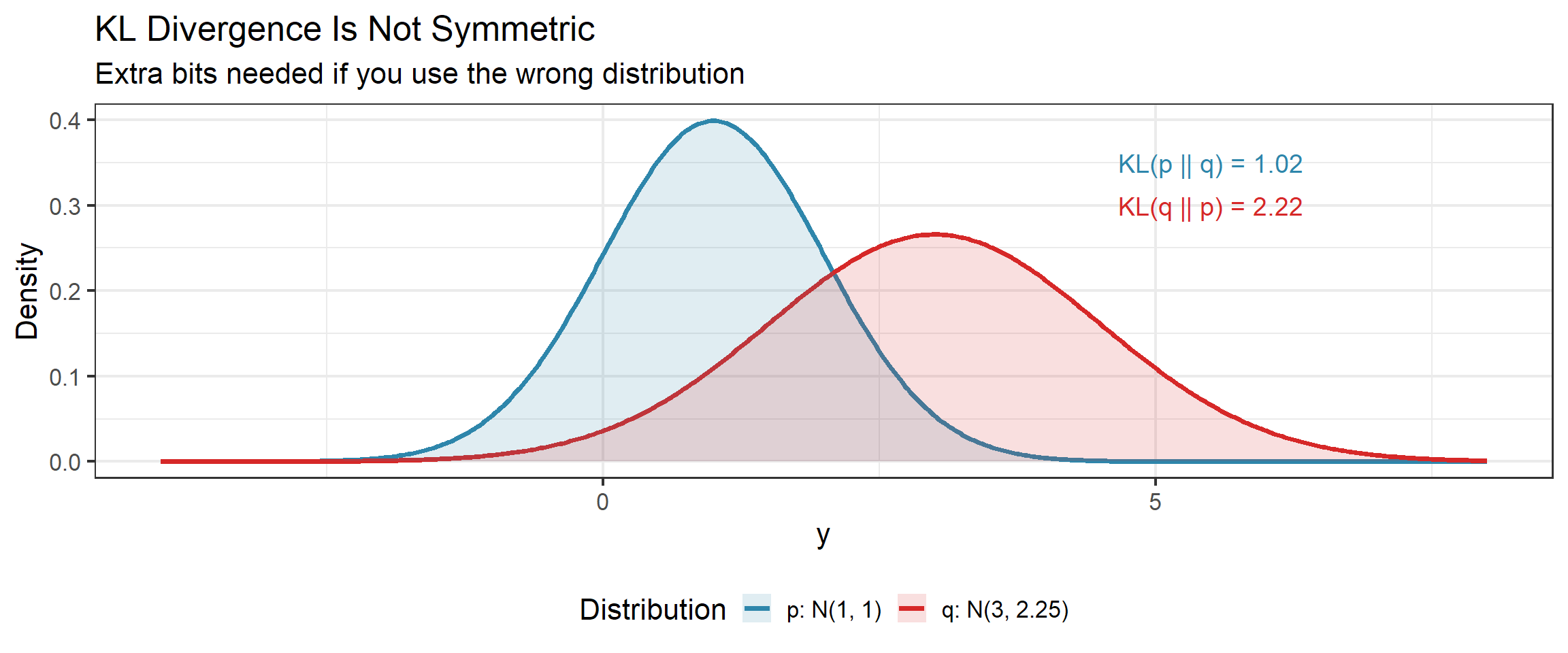
Forward vs Reverse KL: Why Direction Matters
Since \(D_{\text{KL}}(p \,\|\, q) \neq D_{\text{KL}}(q \,\|\, p)\), the two directions have different behavior when we optimize over \(q\):
Forward KL (minimize \(D_{\text{KL}}(p \,\|\, q)\) over \(q\)):
- \(q\) must cover everywhere \(p\) has mass (otherwise \(\log(p/q) \to \infty\))
- Produces mean-seeking approximations — \(q\) spreads to cover all of \(p\)
- This is what MLE does
Reverse KL (minimize \(D_{\text{KL}}(q \,\|\, p)\) over \(q\)):
- \(q\) avoids regions where \(p\) has little mass (since \(q \log(q/p)\) is costly there)
- Produces mode-seeking approximations — \(q\) concentrates on a peak of \(p\)
- This is what variational inference does
A Preview
This asymmetry explains why variational Bayes can underestimate posterior uncertainty: it uses reverse KL, which prefers to lock onto one mode rather than spread across the full posterior. Understanding this distinction is critical for modern Bayesian ML.
KL Divergence: The Likelihood Connection
Here is why KL divergence matters for model selection.
Suppose the truth is \(f\) and our model is \(g_\theta\). We want to find \(\theta\) that minimizes:
\[D_{\text{KL}}(f \,\|\, g_\theta) = E_f\left[\log f(Y)\right] - E_f\left[\log g_\theta(Y)\right]\]
The first term does not depend on \(\theta\). So minimizing KL is equivalent to maximizing:
\[E_f\left[\log g_\theta(Y)\right]\]
This is the expected log-likelihood under the true distribution.
The Key Connection
Minimizing KL divergence \(\iff\) Maximizing expected log-likelihood
This is why maximum likelihood estimation is so natural — it implicitly targets the model closest to truth in KL divergence.
Why KL Divergence Is Fundamental
KL divergence is not just one tool among many — it is the canonical measure of model discrepancy in statistical learning. Here is why it keeps appearing:
| Where it appears | How |
|---|---|
| Maximum likelihood | MLE minimizes \(D_{\text{KL}}(\hat{p}_n \,\|\, q_\theta)\) where \(\hat{p}_n\) is the empirical distribution |
| AIC | Corrects the optimism of MLE as an estimator of expected KL divergence |
| Cross-entropy loss | Training neural nets by minimizing \(-\sum y_i \log \hat{p}_i\) is minimizing KL |
| Variational inference | Approximate posteriors by minimizing reverse KL to the true posterior |
| EM algorithm | Each E-step minimizes KL between the complete-data and incomplete-data posteriors |
| Exponential families | KL between members is the Bregman divergence of the log-partition function |
| Hypothesis testing | Log-likelihood ratio \(= n \cdot D_{\text{KL}}(\hat{p}_n \,\|\, q_0) + o_p(1)\) under the null |
The Unifying Thread
Nearly every major method in statistics and ML can be viewed as minimizing, estimating, or bounding KL divergence. When you see a log-likelihood, you are seeing KL divergence in disguise.
KL Divergence: Summary
| Property | Value |
|---|---|
| Definition | \(E_p[\log(p/q)]\) |
| Decomposition | \(D_{\text{KL}}(p \,\|\, q) = H(p, q) - H(p)\) (cross-entropy minus entropy) |
| Measures | “Extra bits” from using \(q\) when truth is \(p\) |
| Minimum | 0, achieved when \(p = q\) |
| Symmetric? | No — forward KL is mean-seeking, reverse KL is mode-seeking |
| Connection to ML | Minimizing KL \(\iff\) maximizing expected log-likelihood |
| Unit | Nats (base \(e\)) or bits (base 2) |
With this tool in hand, we can now derive AIC.
From KL Divergence to Information Criteria
AIC: The Idea
We want to estimate how well model \(g_\theta\) approximates truth \(f\) in KL divergence.
The natural estimator is the training log-likelihood \(\ell(\hat{\theta})\).
Problem: \(\ell(\hat{\theta})\) is biased upward — the optimism again!
AIC corrects for this optimism.
Under regularity conditions (correct model, large \(n\)), the expected optimism of the log-likelihood is approximately \(d\) (number of free parameters).
Akaike Information Criterion (AIC)
\[\text{AIC} = -2\ell(\hat{\theta}) + 2d\]
Smaller is better. The first term rewards fit; the second penalizes complexity.
Key steps of the derivation:
- Taylor-expand expected log-likelihood around \(\theta_0\)
- The bias of \(\ell(\hat{\theta})\) is approximately \(d\) (Fisher information cancellation)
- Subtract the bias to get AIC
AIC Under Gaussian Errors: Mallow’s Cp
For a Gaussian linear model with known \(\sigma^2\), AIC simplifies beautifully.
Mallow’s Cp
\[C_p = \overline{\text{err}} + \frac{2d \hat{\sigma}^2}{n}\]
where \(\overline{\text{err}}\) is training MSE and \(d\) is the number of parameters.
This is the same covariance penalty we derived earlier!
\[C_p = \text{AIC (Gaussian case)}\]
Intuition
Under Gaussian errors, the optimism of training MSE is exactly \(2d\sigma^2/n\). Mallow’s \(C_p\) simply adds this optimism back. AIC does the same thing, but in log-likelihood units.
BIC: A Different Question
AIC asks: “Which model predicts best?”
BIC asks: “Which model generated the data?”
BIC comes from a Bayesian argument: approximate the marginal likelihood \(p(\mathbf{y} | \mathcal{M})\) via Laplace’s method.
Bayesian Information Criterion (BIC)
\[\text{BIC} = -2\ell(\hat{\theta}) + d \cdot \log(n)\]
Smaller is better.
Key insight: The penalty is \(d \cdot \log(n)\) instead of \(2d\).
Since \(\log(n) > 2\) for \(n \geq 8\), BIC penalizes complexity more heavily than AIC.
AIC vs BIC: Head to Head
| Property | AIC | BIC |
|---|---|---|
| Goal | Best prediction | Identify true model |
| Penalty per parameter | \(2\) | \(\log(n)\) |
| Consistency | No (tends to overfit) | Yes (selects true model as \(n \to \infty\)) |
| Efficiency | Yes (optimal prediction rate) | No |
| Equivalent to… | LOOCV (asymptotically) | Leave-\(n_v\)-out CV where \(n_v/n \to 1\) (Shao, 1997) |
| Favors | Larger models | Smaller models |
Practical Guidance
- Use AIC when prediction accuracy is the goal
- Use BIC when you believe a “true” sparse model exists and want to find it
- When in doubt, report both and see if they agree
Effective Degrees of Freedom
The penalty in AIC/BIC involves \(d\) — the number of free parameters. But for regularized estimators, what does “number of parameters” mean?
Definition: Effective Degrees of Freedom
For a linear smoother \(\hat{\mathbf{y}} = \mathbf{S}\mathbf{y}\):
\[\text{df}(\mathbf{S}) = \text{tr}(\mathbf{S})\]
This is the common currency connecting all the criteria:
- OLS: \(\text{tr}(\mathbf{H}) = p\) (projection matrix has trace = rank)
- Ridge: \(\text{df}(\lambda) = \sum_{j=1}^{p}\frac{d_j^2}{d_j^2 + \lambda}\) where \(d_j\) are singular values of \(\mathbf{X}\)
Ridge df smoothly decreases from \(p\) (at \(\lambda = 0\)) to \(0\) (as \(\lambda \to \infty\)).
Effective df for Ridge: Visualization
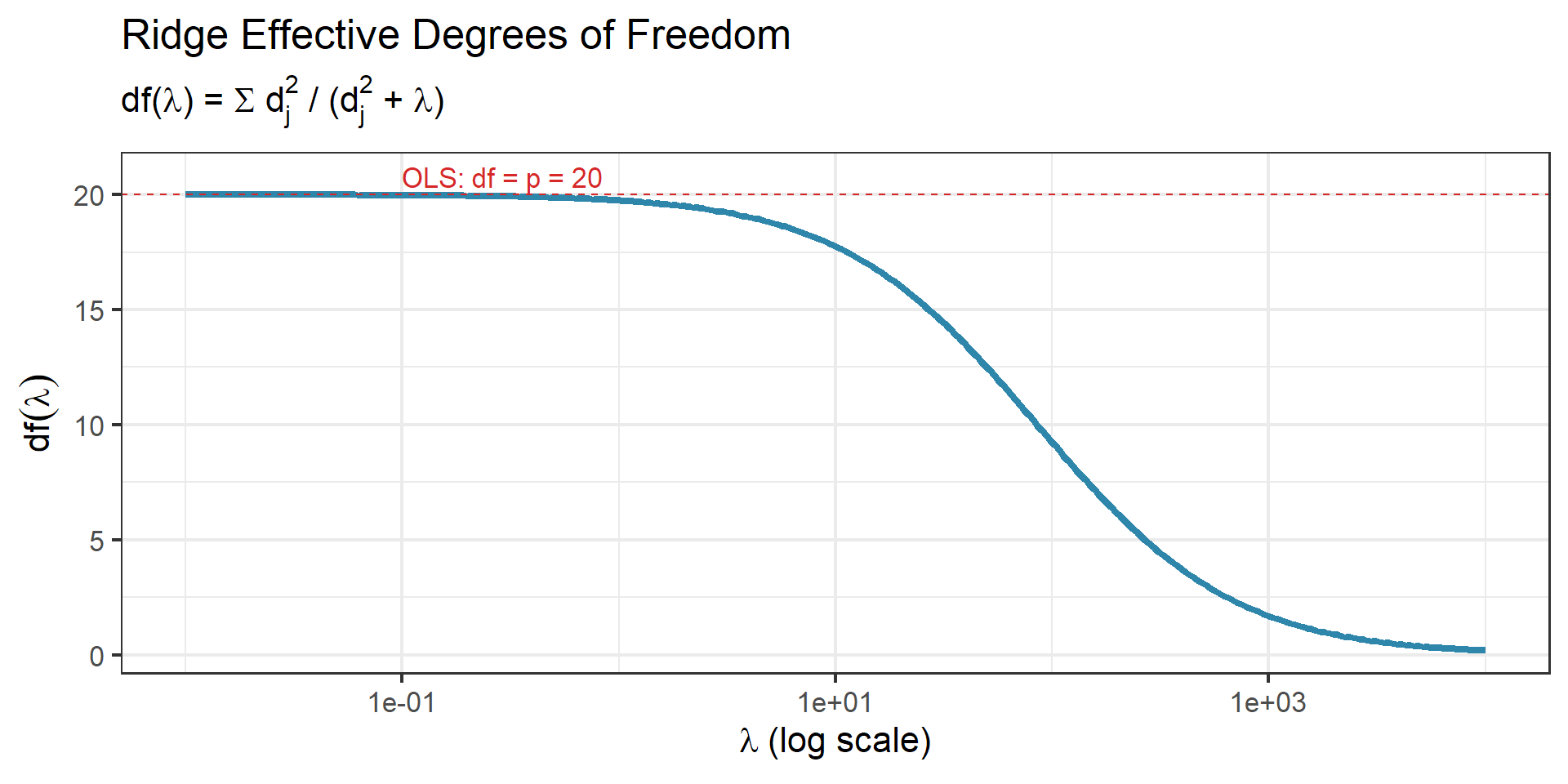
Recall from Week 3
We saw effective df in the context of shrinkage along principal components. Each singular direction contributes \(d_j^2/(d_j^2 + \lambda) \in [0, 1]\) degrees of freedom — a fractional count of how much that direction is being used.
The Stone (1977) Equivalence
A Beautiful Result
AIC \(\approx\) Leave-One-Out Cross-Validation
Stone (1977) showed that for linear models, the model selected by AIC and the model selected by LOOCV are asymptotically identical.
Two completely different philosophies:
- AIC: “Penalize the log-likelihood by \(d\) to correct for optimism” (information theory)
- LOOCV: “Hold out each observation and average prediction errors” (data splitting)
Same answer.
This equivalence is why the information-theoretic and data-splitting camps coexist peacefully. It also motivates the next block: if AIC \(\approx\) LOOCV, we should understand cross-validation deeply.
Cross-Validation
Validation Set Approach
The simplest idea: split data into training and validation sets.
- Fit model on training set
- Evaluate on validation set
Problems:
- High variance: Estimate depends heavily on which observations land in which set
- High bias: Model is trained on only a fraction of the data (often half)
- Wasteful: We discard information
A Poor Estimate
The validation set approach is rarely used on its own. It wastes data and gives unstable estimates. But it introduces the core principle: evaluate on data not used for fitting.
Leave-One-Out Cross-Validation
Idea: Use \(n-1\) observations to train, predict the held-out observation, repeat for all \(n\).
\[\text{CV}(n) = \frac{1}{n}\sum_{i=1}^{n}\left(y_i - \hat{y}_i^{(-i)}\right)^2\]
where \(\hat{y}_i^{(-i)}\) is the prediction for observation \(i\) from the model fit without observation \(i\).
Naive cost: \(n\) model fits. For large datasets, this is prohibitive.
But for linear smoothers, there is a remarkable shortcut.
The Hat Matrix Shortcut
LOOCV via the Hat Matrix
For any linear smoother \(\hat{\mathbf{y}} = \mathbf{S}\mathbf{y}\):
\[\text{CV}(n) = \frac{1}{n}\sum_{i=1}^{n}\left[\frac{y_i - \hat{y}_i}{1 - S_{ii}}\right]^2\]
where \(S_{ii}\) is the \(i\)th diagonal of \(\mathbf{S}\).
Only ONE fit is needed.
Why this works (key insight):
- The residual \(y_i - \hat{y}_i\) underestimates the leave-one-out error
- Dividing by \((1 - S_{ii})\) inflates it by exactly the right amount
- Points with high leverage \(h_i = S_{ii}\) get larger inflation — their residuals are most deflated by having been used in the fit
Connection to Week 2
The hat matrix \(\mathbf{H} = \mathbf{X}(\mathbf{X}^T\mathbf{X})^{-1}\mathbf{X}^T\) keeps returning. The leverages \(h_i = H_{ii}\) measure how much each observation influences its own fit.
\(k\)-Fold Cross-Validation
Divide data into \(k\) roughly equal folds. For each fold \(j\):
- Train on all folds except \(j\)
- Compute MSE on fold \(j\)
\[\text{CV}(k) = \frac{1}{k}\sum_{j=1}^{k}\text{MSE}_j\]
The bias-variance tradeoff in \(k\):
| \(k\) | Bias | Variance | Computation |
|---|---|---|---|
| \(n\) (LOOCV) | Low | High (correlated folds) | \(n\) fits |
| 10 | Slightly higher | Lower | 10 fits |
| 5 | Moderate | Lowest | 5 fits |
Why does LOOCV have high variance? Each of the \(n\) training sets shares \(n-2\) observations with every other. Averaging highly correlated quantities does not reduce variance much.
The Standard Choice
\(k = 5\) or \(k = 10\) is nearly universally recommended (Breiman & Spector, 1992; Hastie et al., 2009). The slight bias increase is more than compensated by variance reduction.
The 1-SE rule (Breiman et al., 1984): Select the most parsimonious model whose CV error is within one standard error of the minimum. This trades a negligible increase in error for a simpler model. cv.glmnet uses this as default (lambda.1se).
Generalized Cross-Validation (GCV)
For linear smoothers, replace each individual leverage \(S_{ii}\) with the average leverage \(\text{tr}(\mathbf{S})/n\):
GCV Formula
\[\text{GCV} = \frac{1}{n}\sum_{i=1}^{n}\left[\frac{y_i - \hat{f}(x_i)}{1 - \text{tr}(\mathbf{S})/n}\right]^2\]
Advantages:
- No need to compute individual leverages
- For Ridge: only need \(\text{df}(\lambda) = \text{tr}(\mathbf{S}_\lambda)\), which has a closed form
- Computationally cheap, good approximation to LOOCV
This bridges Weeks 3 and 4: the effective degrees of freedom from Ridge regression now plug directly into GCV for model selection.
The Chain of Equivalences
The Unifying Chain
\[C_p = \text{AIC (Gaussian)} \;\approx\; \text{LOOCV (Stone, 1977)} \;\approx\; \text{GCV (linear smoothers)}\]
All correct for the same thing: the optimism of training error.
Four different derivations:
- \(C_p\): covariance penalty
- AIC: KL divergence + bias correction
- LOOCV: data splitting
- GCV: average-leverage approximation to LOOCV
Why This Matters
When theorists from different schools converge on the same answer, it is a strong signal that the answer is right. The optimism of training error really is \(2d\sigma^2/n\) for linear smoothers, no matter how you approach the problem.
The Wrong Way to Do CV
A Catastrophic Mistake (ESL 7.10.2)
WRONG: Screen features on all data, then cross-validate on the selected features.
Example from ESL:
- 50 observations, 5000 features, true error rate = 50% (pure noise)
- Screen to top 100 features using all data
- 5-fold CV on the 100 selected features reports 3% error
The CV thinks we have a great classifier. In reality, we have nothing.
The fix is simple:
The Right Way
All supervised steps must go INSIDE the CV loop.
Feature screening, variable selection, hyperparameter tuning — everything that looks at \(Y\) must be repeated within each fold.
Connection to Modern ML Practice
In production ML pipelines, this principle extends to nested cross-validation: an outer loop estimates generalization error, while an inner loop tunes hyperparameters (learning rate, regularization, architecture). Grid search, random search, and Bayesian optimization (e.g., scikit-learn’s GridSearchCV) all implement this internally. The principle is unchanged from what we see here — only the scale differs.
If you remember one thing from this slide, let it be this.
Code Demo: k-Fold CV from Scratch
Code Demo: k-Fold CV with glmnet
set.seed(577)
n <- 200; p <- 50
X <- matrix(rnorm(n * p), n, p)
beta_true <- c(rep(2, 5), rep(0, 45))
y <- X %*% beta_true + rnorm(n, sd = 2)
# 10-fold CV for Lasso
cv_fit <- cv.glmnet(X, y, alpha = 1, nfolds = 10)
# Rebuild CV curve in ggplot2 for consistent styling
cv_df <- tibble(
lambda = cv_fit$lambda,
cvm = cv_fit$cvm,
cvup = cv_fit$cvup,
cvlo = cv_fit$cvlo,
nzero = cv_fit$nzero
)
p_cv <- ggplot(cv_df, aes(x = log(lambda), y = cvm)) +
geom_ribbon(aes(ymin = cvlo, ymax = cvup), alpha = 0.15, fill = colors["blue"]) +
geom_point(color = colors["red"], size = 1.8) +
geom_vline(xintercept = log(cv_fit$lambda.min), linetype = "dashed", color = colors["blue"]) +
geom_vline(xintercept = log(cv_fit$lambda.1se), linetype = "dashed", color = colors["orange"]) +
annotate("text", x = log(cv_fit$lambda.min), y = max(cv_df$cvup) * 0.95,
label = "lambda[min]", parse = TRUE, hjust = 1.1, color = colors["blue"], size = 4.5) +
annotate("text", x = log(cv_fit$lambda.1se), y = max(cv_df$cvup) * 0.95,
label = "lambda['1se']", parse = TRUE, hjust = -0.1, color = colors["orange"], size = 4.5) +
labs(title = "10-Fold Cross-Validation for Lasso",
subtitle = "Error bars = +/- 1 SE | Dashed lines = lambda_min and lambda_1se",
x = expression(log(lambda)), y = "Mean Squared Error (CV)") +
theme_bw(base_size = 16) +
theme(plot.title = element_text(size = 14))
ggsave(file.path(fig_dir, "05_cv_glmnet.png"), p_cv, width = 10, height = 5, dpi = 150)
p_cv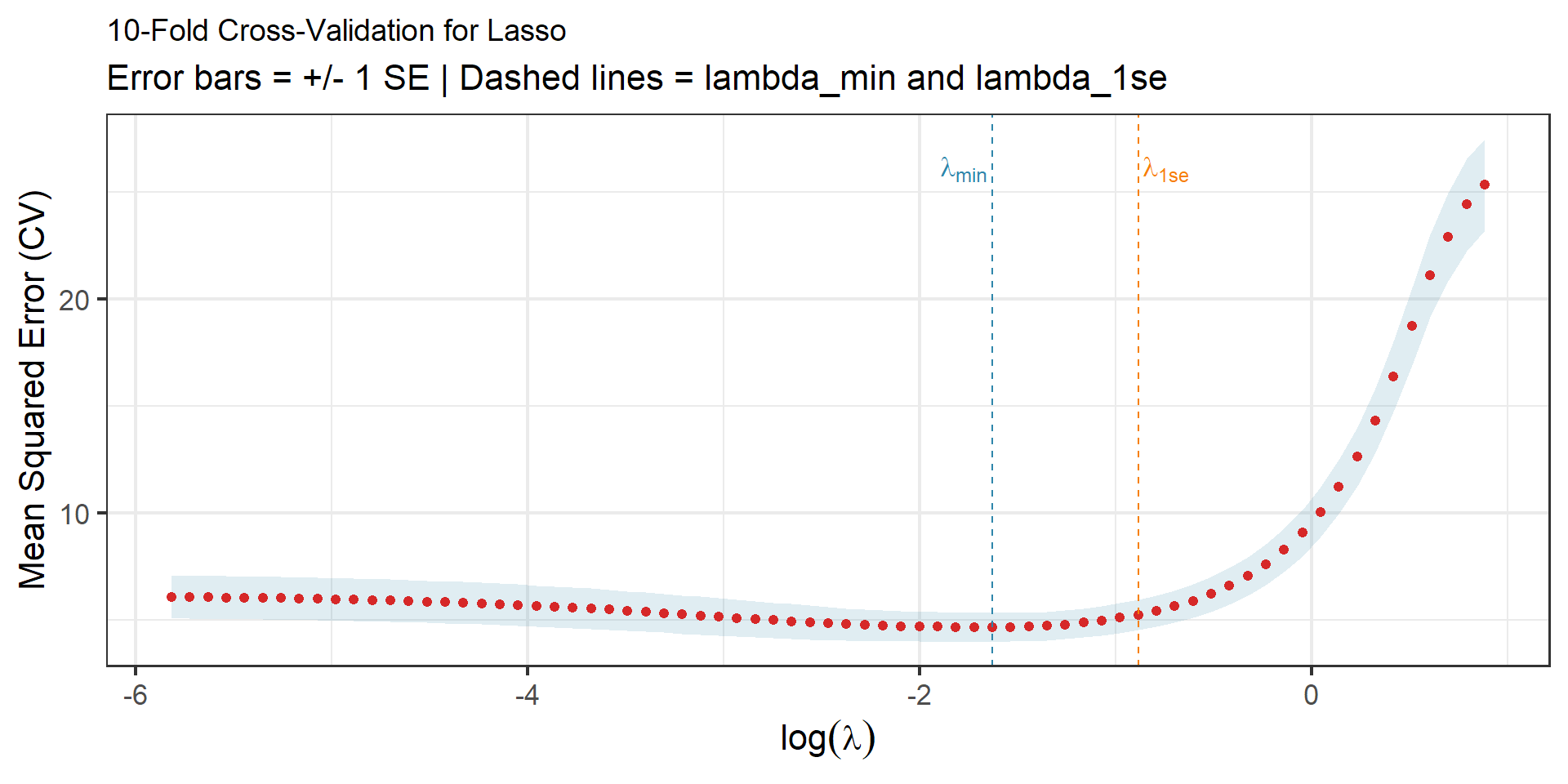
Reading the Plot
- Left dashed line: \(\lambda_{\min}\) — minimizes CV error
- Right dashed line: \(\lambda_{\text{1se}}\) — most regularized model within 1 SE of the minimum
- The x-axis is \(\log(\lambda)\), so regularization increases from left to right
Bootstrap
The Bootstrap Idea
Efron (1979): Instead of deriving the sampling distribution analytically, simulate it.
Algorithm:
- Draw a sample of size \(n\) with replacement from the data
- Compute your statistic \(\hat{\theta}^*_b\) on this resample
- Repeat \(B\) times
- Use the empirical distribution of \(\{\hat{\theta}^*_1, \ldots, \hat{\theta}^*_B\}\) to estimate variability
\[\widehat{\text{SE}}_B = \sqrt{\frac{1}{B-1}\sum_{b=1}^{B}\left(\hat{\theta}^*_b - \bar{\theta}^*\right)^2}\]
The Power of Bootstrap
Works for any statistic — medians, quantiles, ratios, eigenvalues, anything. No formulas needed for the sampling distribution.
Why 63.2% Matters
What fraction of the original data appears in a bootstrap sample?
\[P(\text{observation } i \text{ not selected in any of } n \text{ draws}) = \left(1 - \frac{1}{n}\right)^n \to e^{-1} \approx 0.368\]
So each bootstrap sample contains \(\approx 63.2\%\) unique observations.
The remaining \(\approx 36.8\%\) are left out — they form a free validation set.
A Recurring Idea
This 63.2/36.8 split is not just a curiosity. It is the basis of:
- The .632 bootstrap estimator for prediction error
- Out-of-bag (OOB) error in random forests (Week 6)
- Various bagging strategies for variance reduction
Bootstrap for Prediction Error
Naive bootstrap: Compute test error of each bootstrap model on all \(n\) observations.
Problem: Training and test sets overlap — the estimate is overly optimistic.
The .632 estimator corrects this:
\[\widehat{\text{Err}}_{.632} = 0.368 \cdot \overline{\text{err}} + 0.632 \cdot \widehat{\text{Err}}_1\]
where \(\widehat{\text{Err}}_1\) is the average error on left-out observations across all bootstrap samples.
The .632+ estimator adapts the weights when overfitting is severe — it adjusts the mixing proportion based on the degree of overfitting (no-information error rate).
Preview: Out-of-Bag Error
The 36.8% of observations left out of each bootstrap sample give us free validation data.
For random forests (Week 6):
- Each tree is trained on a bootstrap sample
- For observation \(i\), average predictions only from trees where \(i\) was left out
- This gives the OOB error — no separate validation set needed
Looking Ahead
We will see this idea return as the primary error estimate for bagged and random forest models. The bootstrap is not just a theoretical tool — it is baked into the machinery of ensemble methods.
Bootstrap Demo: Ridge Coefficient Uncertainty
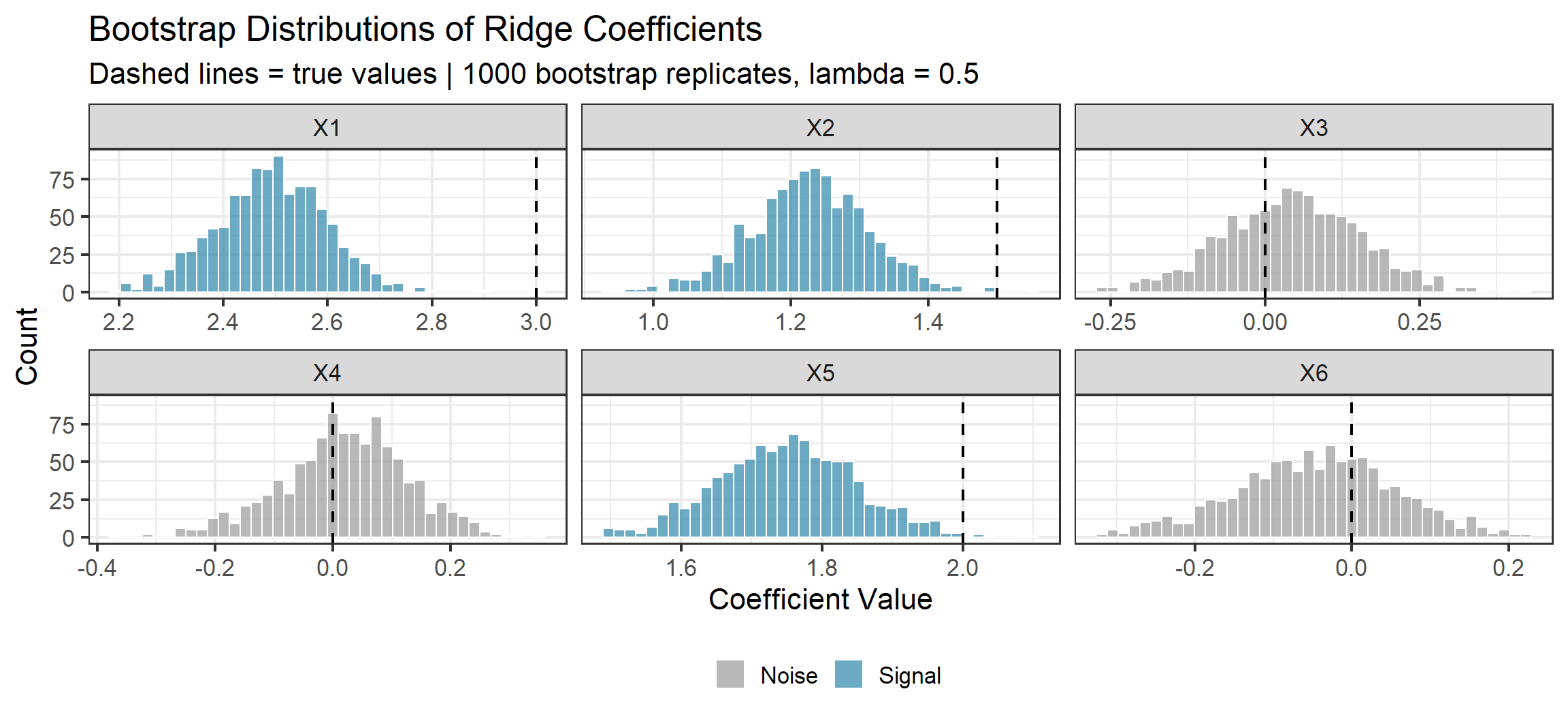
Notice the Shrinkage
The bootstrap distributions for the signal coefficients (\(X_1, X_2, X_5\)) are centered below their true values. This is Ridge doing its job — biasing toward zero in exchange for lower variance.
Bootstrap Confidence Intervals
Beyond standard errors, the bootstrap gives us confidence intervals without distributional assumptions.
Three main approaches:
| Method | Interval | Pros/Cons |
|---|---|---|
| Percentile | \([\hat{\theta}^*_{\alpha/2},\; \hat{\theta}^*_{1-\alpha/2}]\) | Simple; can have poor coverage for skewed distributions |
| Basic (pivotal) | \([2\hat{\theta} - \hat{\theta}^*_{1-\alpha/2},\; 2\hat{\theta} - \hat{\theta}^*_{\alpha/2}]\) | Uses symmetry of \(\hat{\theta}^* - \hat{\theta}\) around 0 |
| BCa | Bias-corrected and accelerated | Best coverage; adjusts for bias and skewness |
Practical Recommendation
Use BCa (bias-corrected and accelerated) when possible. It adjusts for both bias in the bootstrap distribution and skewness, giving second-order accurate coverage: the coverage error is \(O(1/n)\) rather than \(O(1/\sqrt{n})\) for the percentile method.
Bootstrap CI Demo: Median Regression Coefficient
set.seed(577)
n <- 100; p <- 10
X <- matrix(rnorm(n * p), n, p)
beta_true <- c(3, 1.5, 0, 0, 2, rep(0, 5))
y <- X %*% beta_true + rnorm(n)
# Point estimate: Ridge coefficient for X1
fit_full <- glmnet(X, y, alpha = 0, lambda = 0.5)
theta_hat <- as.vector(coef(fit_full))[-1][1] # X1 coefficient
# Bootstrap
B <- 2000
boot_theta <- numeric(B)
for (b in 1:B) {
idx <- sample(1:n, n, replace = TRUE)
fit_b <- glmnet(X[idx, ], y[idx], alpha = 0, lambda = 0.5)
boot_theta[b] <- as.vector(coef(fit_b))[-1][1]
}
# Three types of 95% CIs
alpha <- 0.05
q_lo <- quantile(boot_theta, alpha / 2)
q_hi <- quantile(boot_theta, 1 - alpha / 2)
ci_percentile <- c(q_lo, q_hi)
ci_basic <- c(2 * theta_hat - q_hi, 2 * theta_hat - q_lo)
ci_df <- tibble(
Method = c("Percentile", "Basic (Pivotal)"),
lo = c(ci_percentile[1], ci_basic[1]),
hi = c(ci_percentile[2], ci_basic[2]),
ypos = c(-0.15, 0)
)
p_ci <- ggplot() +
geom_histogram(aes(x = boot_theta, y = after_stat(density)),
bins = 50, fill = colors["blue"], alpha = 0.5, color = "white") +
geom_vline(xintercept = theta_hat, linewidth = 1, color = "black") +
geom_vline(xintercept = 3, linewidth = 1, linetype = "dashed", color = colors["red"]) +
geom_segment(data = ci_df, aes(x = lo, xend = hi, y = ypos, yend = ypos, color = Method),
linewidth = 2) +
geom_point(data = ci_df, aes(x = lo, y = ypos, color = Method), size = 3) +
geom_point(data = ci_df, aes(x = hi, y = ypos, color = Method), size = 3) +
scale_color_manual(values = c("Percentile" = colors["green"], "Basic (Pivotal)" = colors["orange"])) +
annotate("text", x = 3.05, y = 2.5, label = "'True' ~ beta[1] == 3",
parse = TRUE, color = colors["red"], hjust = 0, size = 4.5) +
annotate("text", x = theta_hat + 0.05, y = 2.8,
label = paste0("hat(theta) == ", round(theta_hat, 2)),
parse = TRUE, hjust = 0, size = 4.5) +
labs(title = "Bootstrap Confidence Intervals for Ridge Coefficient (X1)",
subtitle = "2000 bootstrap replicates | lambda = 0.5 | 95% confidence level",
x = "Coefficient Value", y = "Density") +
theme_bw(base_size = 16) +
theme(legend.position = "bottom")
ggsave(file.path(fig_dir, "06_bootstrap_ci.png"), p_ci, width = 10, height = 5, dpi = 150)
p_ci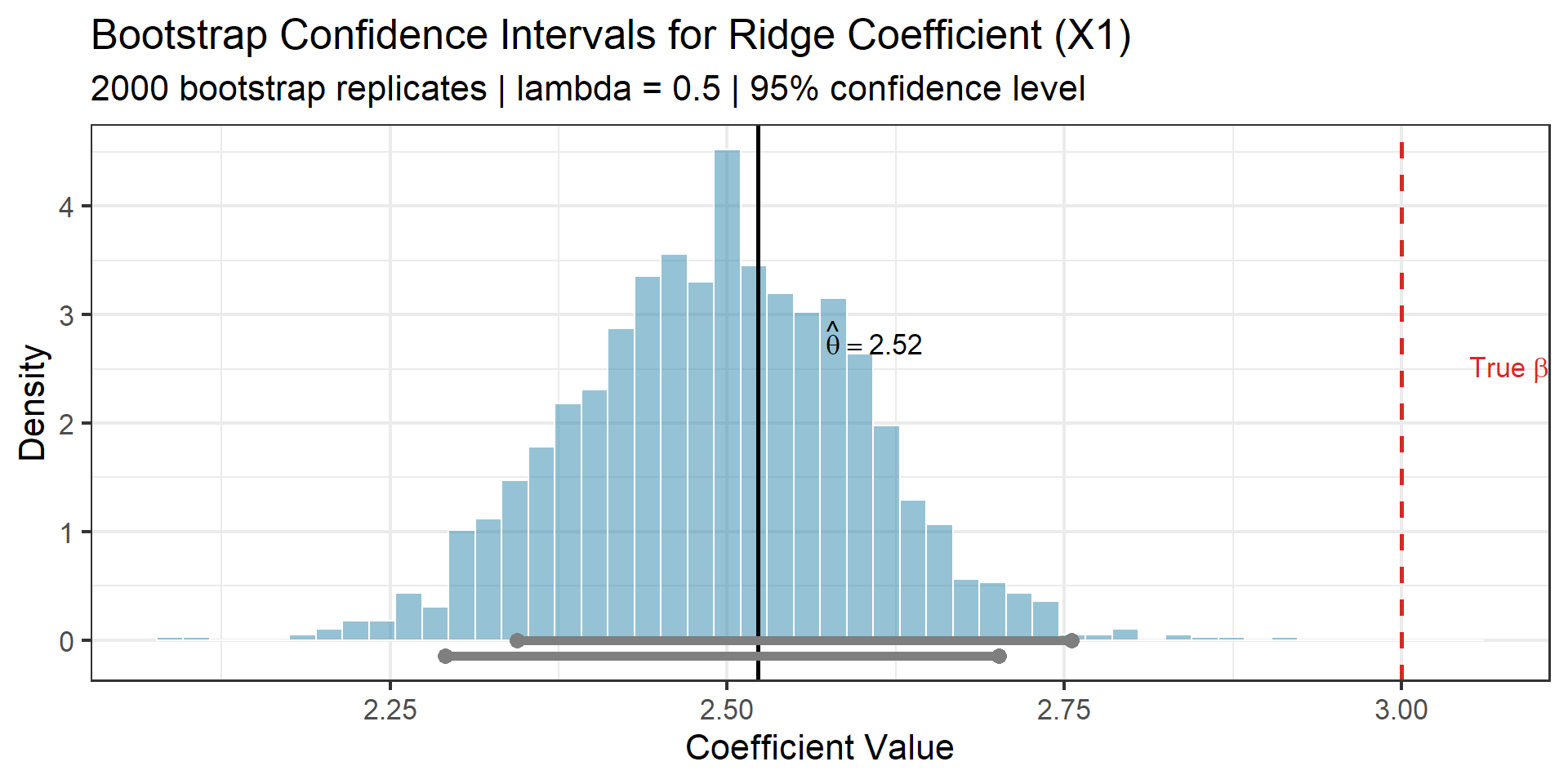
Observe
Both intervals contain the true value \(\beta_1 = 3\), but the Ridge point estimate \(\hat{\theta}\) is biased below it. The bootstrap honestly reflects this bias — the distribution is centered at the biased estimate, not the truth. This is a feature, not a bug: bootstrap intervals for shrinkage estimators correctly account for the bias-variance trade-off.
Bootstrap → Bagging
Bagging (Bootstrap AGGregatING) is literally the bootstrap applied to prediction: fit \(B\) models on bootstrap samples, average predictions. This reduces variance by a factor related to the correlation between bootstrap replicates. We will make this precise in Week 6.
Conformal Prediction
The Problem with Classical Prediction Intervals
Classical prediction intervals assume a model:
\[Y = \mathbf{x}^T\boldsymbol{\beta} + \varepsilon, \qquad \varepsilon \sim N(0, \sigma^2)\]
What if normality fails? What if we are using a random forest, or a neural network?
We need prediction intervals for black-box models with finite-sample guarantees.
Conformal Prediction: The Promise
Given any model and any distribution (subject to one mild assumption), conformal prediction delivers intervals with guaranteed coverage — in finite samples, not just asymptotically.
Conformal prediction is increasingly adopted in high-stakes ML applications — medical diagnostics, autonomous driving, drug discovery — where regulatory requirements demand quantified uncertainty, not just point predictions.
The Only Assumption: Exchangeability
Definition: Exchangeability
Random variables \((Z_1, \ldots, Z_n, Z_{n+1})\) are exchangeable if their joint distribution is invariant under all permutations:
\[(Z_1, \ldots, Z_{n+1}) \stackrel{d}{=} (Z_{\pi(1)}, \ldots, Z_{\pi(n+1)})\]
for every permutation \(\pi\).
i.i.d. \(\implies\) exchangeable (but exchangeability is weaker).
When exchangeability fails:
- Time series: observations have temporal order
- Adaptive data collection: later samples depend on earlier ones
- Distribution shift: training and test distributions differ
Split Conformal: The Algorithm
Split Conformal Prediction
Input: Data \((X_1, Y_1), \ldots, (X_n, Y_n)\), new point \(X_{n+1}\), level \(\alpha\)
- Split data into training \(\mathcal{I}_1\) and calibration \(\mathcal{I}_2\)
- Fit any model \(\hat{f}\) on \(\mathcal{I}_1\)
- Score: Compute residuals \(R_i = |Y_i - \hat{f}(X_i)|\) for \(i \in \mathcal{I}_2\)
- Quantile: Let \(\hat{q}\) be the \(\lceil(1-\alpha)(|\mathcal{I}_2|+1)\rceil\)-th smallest value among \(\{R_i : i \in \mathcal{I}_2\}\)
- Interval: \(C(X_{n+1}) = [\hat{f}(X_{n+1}) - \hat{q},\; \hat{f}(X_{n+1}) + \hat{q}]\)
That is the entire algorithm. Five lines. Model-agnostic.
The Coverage Guarantee
Finite-Sample Coverage
Under exchangeability:
\[1 - \alpha \;\leq\; P\!\left(Y_{n+1} \in C(X_{n+1})\right) \;\leq\; 1 - \alpha + \frac{1}{|\mathcal{I}_2| + 1}\]
This is remarkable:
- Finite-sample — not asymptotic
- Distribution-free — no parametric assumptions
- Model-agnostic — works with linear regression, random forests, neural networks, anything
Proof intuition: Under exchangeability, the test residual \(R_{n+1}\) has a rank that is uniformly distributed among the \(|\mathcal{I}_2| + 1\) residuals. The probability that it exceeds the \((1-\alpha)\)-quantile is at most \(\alpha\).
Coverage Is Free, Width Is Earned
What conformal gives for free:
- Valid coverage at level \(1 - \alpha\)
- This works for any model
- Even a terrible model gets valid coverage
What you must earn:
- Narrow intervals
- A better model \(\implies\) smaller residuals \(\implies\) smaller \(\hat{q}\) \(\implies\) tighter intervals
- The score function matters too
Score choices:
| Score | Intervals | Best when |
|---|---|---|
| \(|Y - \hat{f}(X)|\) | Constant width | Homoskedastic data |
| \(|Y - \hat{f}(X)| / \hat{\sigma}(X)\) | Adaptive width | Heteroskedastic data |
Conformal vs Other Prediction Intervals
| Property | Conformal | Bootstrap | Parametric |
|---|---|---|---|
| Assumption | Exchangeability | Smooth \(F\) | Normality |
| Coverage | Finite-sample | Asymptotic | Exact if model correct |
| Computational cost | 1 fit | \(B\) fits | 1 fit |
| Model-agnostic? | Yes | Yes | No |
| Adaptive width? | With normalized scores | Yes | Yes |
Conformal for Classification
Replace prediction intervals with prediction sets:
- Define score: \(s(x, y) = 1 - \hat{\pi}_y(x)\) (one minus predicted probability of true class)
- Compute calibration scores: \(R_i = s(X_i, Y_i)\) for \(i \in \mathcal{I}_2\)
- Quantile: \(\hat{q}\) as before
- Prediction set: \(C(X_{n+1}) = \{y : \hat{\pi}_y(X_{n+1}) \geq 1 - \hat{q}\}\)
Interpretation: Include all classes whose predicted probability exceeds a threshold calibrated to guarantee coverage.
What Prediction Sets Tell You
- A confident, correct model produces singleton sets (one class)
- An uncertain model produces larger sets (multiple classes)
- A terrible model includes almost all classes — coverage is free, precision is earned
Conformal Demo: Heteroskedastic Data
set.seed(577)
n <- 500
x <- runif(n, 0, 5)
y <- sin(2 * x) + rnorm(n, sd = 0.2 + 0.3 * x)
# Split into fit / calibrate / test
fit_idx <- 1:200; cal_idx <- 201:400; test_idx <- 401:500
# Fit model on training
df_fit <- data.frame(x = x[fit_idx], y = y[fit_idx])
fit <- lm(y ~ poly(x, 5), data = df_fit)
# Calibration scores
cal_pred <- predict(fit, newdata = data.frame(x = x[cal_idx]))
cal_scores <- abs(y[cal_idx] - cal_pred)
# Conformal quantile (90% coverage)
alpha <- 0.10
n_cal <- length(cal_idx)
q_level <- ceiling((1 - alpha) * (n_cal + 1)) / n_cal
qhat <- quantile(cal_scores, probs = min(q_level, 1), type = 1)
# Test predictions and intervals
test_pred <- predict(fit, newdata = data.frame(x = x[test_idx]))
# Also compute parametric intervals
pred_se <- predict(fit, newdata = data.frame(x = x[test_idx]), se.fit = TRUE)
param_margin <- qt(1 - alpha/2, df = fit$df.residual) *
sqrt(pred_se$se.fit^2 + summary(fit)$sigma^2)
# Empirical coverage
conformal_coverage <- mean(y[test_idx] >= test_pred - qhat &
y[test_idx] <= test_pred + qhat)
param_coverage <- mean(y[test_idx] >= test_pred - param_margin &
y[test_idx] <= test_pred + param_margin)
test_df <- tibble(
x = x[test_idx], y = y[test_idx],
pred = test_pred,
conf_lo = test_pred - qhat, conf_hi = test_pred + qhat,
param_lo = test_pred - param_margin, param_hi = test_pred + param_margin
)
p_plot <- ggplot(test_df, aes(x = x)) +
# Conformal bands
geom_ribbon(aes(ymin = conf_lo, ymax = conf_hi), fill = colors["blue"], alpha = 0.25) +
# Parametric bands
geom_ribbon(aes(ymin = param_lo, ymax = param_hi), fill = colors["red"], alpha = 0.15) +
# Data and prediction
geom_point(aes(y = y), size = 1.5, alpha = 0.6) +
geom_line(aes(y = pred), color = "black", linewidth = 1) +
annotate("text", x = 0.3, y = 3.5,
label = paste0("Conformal (", round(100*conformal_coverage), "% coverage)"),
color = colors["blue"], size = 4.5, hjust = 0, fontface = "bold") +
annotate("text", x = 0.3, y = 3.0,
label = paste0("Parametric (", round(100*param_coverage), "% coverage)"),
color = colors["red"], size = 4.5, hjust = 0, fontface = "bold") +
labs(title = "Split Conformal vs Parametric Prediction Intervals",
subtitle = "Heteroskedastic data: variance increases with x | Target: 90% coverage",
x = "x", y = "y") +
theme_bw(base_size = 16)
ggsave(file.path(fig_dir, "04_conformal_demo.png"), p_plot, width = 12, height = 5.5, dpi = 150)
p_plot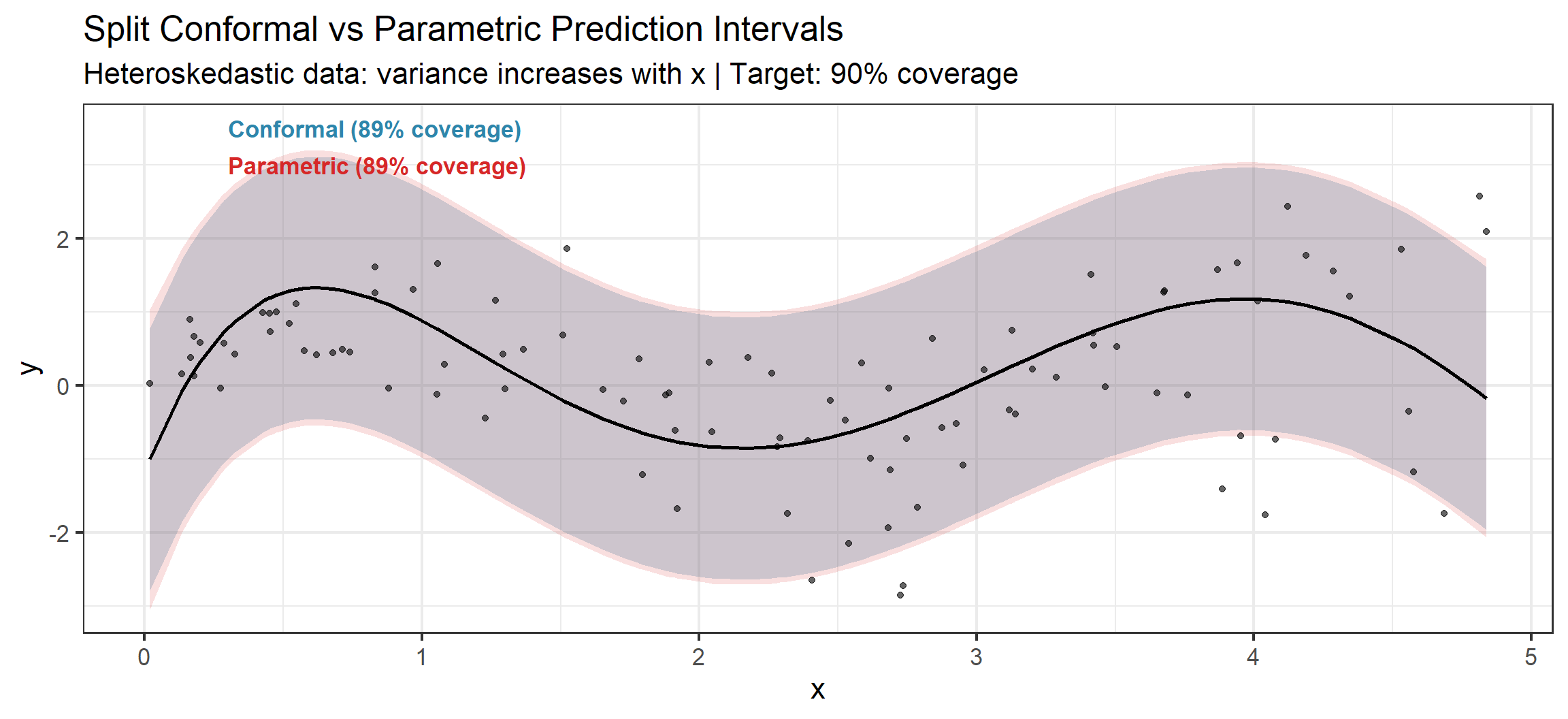
Notice the Difference
Conformal bands have constant width (since we used absolute residual scores). They are too wide on the left and too narrow on the right. Using normalized scores \(|Y - \hat{f}(X)| / \hat{\sigma}(X)\) would produce adaptive-width bands that better match the heteroskedasticity.
Synthesis
The Chain Revisited
\[C_p = \text{AIC (Gaussian)} \;\approx\; \text{LOOCV (Stone, 1977)} \;\approx\; \text{GCV (linear smoothers)}\]
Four tools, four different stories:
| Tool | What it tells you |
|---|---|
| CV | “Here is an estimate of the error rate” |
| Bootstrap | “Here is the uncertainty in our estimates” |
| AIC / BIC | “Here is the analytically optimal penalty” |
| Conformal | “Here is a guaranteed prediction set” |
When to Use What
| Situation | Recommended Method |
|---|---|
| Comparing a few nested linear models | AIC or BIC |
| Tuning \(\lambda\) for Ridge/Lasso | \(k\)-fold CV (\(k = 5\) or \(10\)) |
| Estimating SE of any statistic | Bootstrap |
| Black-box model, need prediction intervals | Conformal prediction |
| Linear smoother, quick approximation | GCV |
| Believe a true sparse model exists | BIC |
| Best predictive performance | AIC or CV |
The Week 4 Narrative
We started by asking how to choose \(\lambda\) and ended up connecting information theory, Bayesian model evidence, data splitting, and distribution-free inference. These are not competing approaches — they are complementary views of the same fundamental problem: how to honestly assess a model’s performance.
Week 5 Synthesis: Resampling vs Conformal
The Same Question, Two Philosophies
We have spent this week building tools to honestly assess model performance. But resampling and conformal prediction answer this question in fundamentally different ways.
The Core Distinction
Resampling (bootstrap, CV) asks: “How variable are my estimates?”
Conformal prediction asks: “Where will the next observation fall?”
This is not a difference in technique — it is a difference in what you are trying to guarantee.
- Resampling targets parameters and model performance (estimation)
- Conformal targets individual predictions (prediction)
What Each Framework Gives You
Resampling Methods
Bootstrap, Cross-Validation, Jackknife
- Standard errors of \(\hat{\beta}\), \(\hat{\theta}\)
- Confidence intervals for parameters
- Estimates of test error (CV)
- Model selection (which \(\lambda\)?)
- Bias correction (\(C_p\), AIC, GCV)
These are statements about the model or the procedure.
Conformal Prediction
Split conformal, CQR, Jackknife+
- Prediction intervals for \(Y_{n+1}\)
- Prediction sets for classification
- Coverage guarantees at level \(1 - \alpha\)
- Uncertainty quantification per observation
These are statements about the next data point.
A Useful Litmus Test
Ask yourself: Am I trying to learn about the model, or promise something about a future observation? Your answer determines which framework you need.
The Guarantee Gap
The deepest difference is in what each framework promises and what it assumes to promise it.
| Resampling | Conformal | |
|---|---|---|
| Type of guarantee | Asymptotic (usually) | Finite-sample |
| Key assumption | Smooth \(F\), regularity conditions, or correct model | Exchangeability |
| What can break it | Small \(n\), model misspecification, non-smooth functionals | Distribution shift, non-exchangeability |
| Coverage | Approximate: \(P(\theta \in \text{CI}) \approx 1 - \alpha\) | Finite-sample valid: \(P(Y_{n+1} \in C) \geq 1 - \alpha\) |
Why this matters in practice:
- A bootstrap 95% CI might have 88% actual coverage if \(n\) is small or the statistic is non-smooth
- A conformal 95% interval has \(\geq 95\)% coverage by construction, regardless of \(n\) or model quality (though a bad model yields uninformatively wide intervals)
The price conformal pays: it says nothing about the model’s parameters. You get a prediction set, not understanding.
Shared Machinery, Different Goals
Despite different philosophies, the mechanics often overlap:
Residuals are the common language.
- CV computes residuals \(y_i - \hat{f}^{(-i)}(x_i)\) to estimate test error
- Conformal computes residuals \(|y_i - \hat{f}_{\mathcal{I}_1}(x_i)|\) on calibration data to set prediction intervals
- Bootstrap resamples residuals to approximate sampling distributions
Data splitting appears everywhere.
- \(k\)-fold CV splits data into \(k\) folds for error estimation
- Split conformal splits data into training and calibration sets
- Out-of-bag (OOB) in bagging is a bootstrap-based split
The Jackknife+ Bridge
Barber et al. (2021) showed that LOO residuals — the bread and butter of cross-validation — can be repurposed to give distribution-free coverage guarantees (the jackknife+, with coverage at level \(1 - 2\alpha\)). Same residuals, upgraded guarantee. This is the clearest example of resampling machinery serving conformal goals.
What Conformal Cannot Do (and Resampling Can)
Conformal prediction is powerful, but it has blind spots:
1. No parameter inference. You cannot use conformal to build a confidence interval for \(\beta_1\) in a linear model. That requires the bootstrap or classical theory.
2. No model comparison. Conformal tells you the width of your prediction interval, but it does not tell you which model to pick. You still need CV or AIC/BIC for that.
3. No standard errors. If someone asks “how stable is your estimate?”, conformal has no answer. Bootstrap does.
4. Marginal coverage only. The guarantee \(P(Y_{n+1} \in C(X_{n+1})) \geq 1 - \alpha\) is marginal over the randomness of the entire dataset. It does not guarantee coverage conditionally on \(X_{n+1} = x\). Exact conditional coverage is provably impossible without further assumptions (Lei & Wasserman, 2014), though approximate and group-conditional variants are an active research area.
What Resampling Cannot Do (and Conformal Can)
1. Finite-sample guarantees. Bootstrap CIs rely on \(n \to \infty\). Conformal coverage holds for \(n = 50\) just as well as \(n = 50{,}000\).
2. True model-agnosticism. Bootstrap intervals for a neural network require careful thought about what “resample” means in that context. Conformal treats \(\hat{f}\) as a black box — any algorithm, any complexity.
3. Robustness to model misspecification. If your model is wrong, bootstrap CIs can have poor coverage. Conformal coverage holds regardless — you just get wider intervals.
A Subtle Point
The bootstrap can be applied to any statistic, but its theoretical guarantees degrade for non-smooth functionals (e.g., quantiles, max). Conformal makes no smoothness assumptions — its coverage guarantee is permutation-based (exchangeability of ranks), not analytic.
The Decision Framework
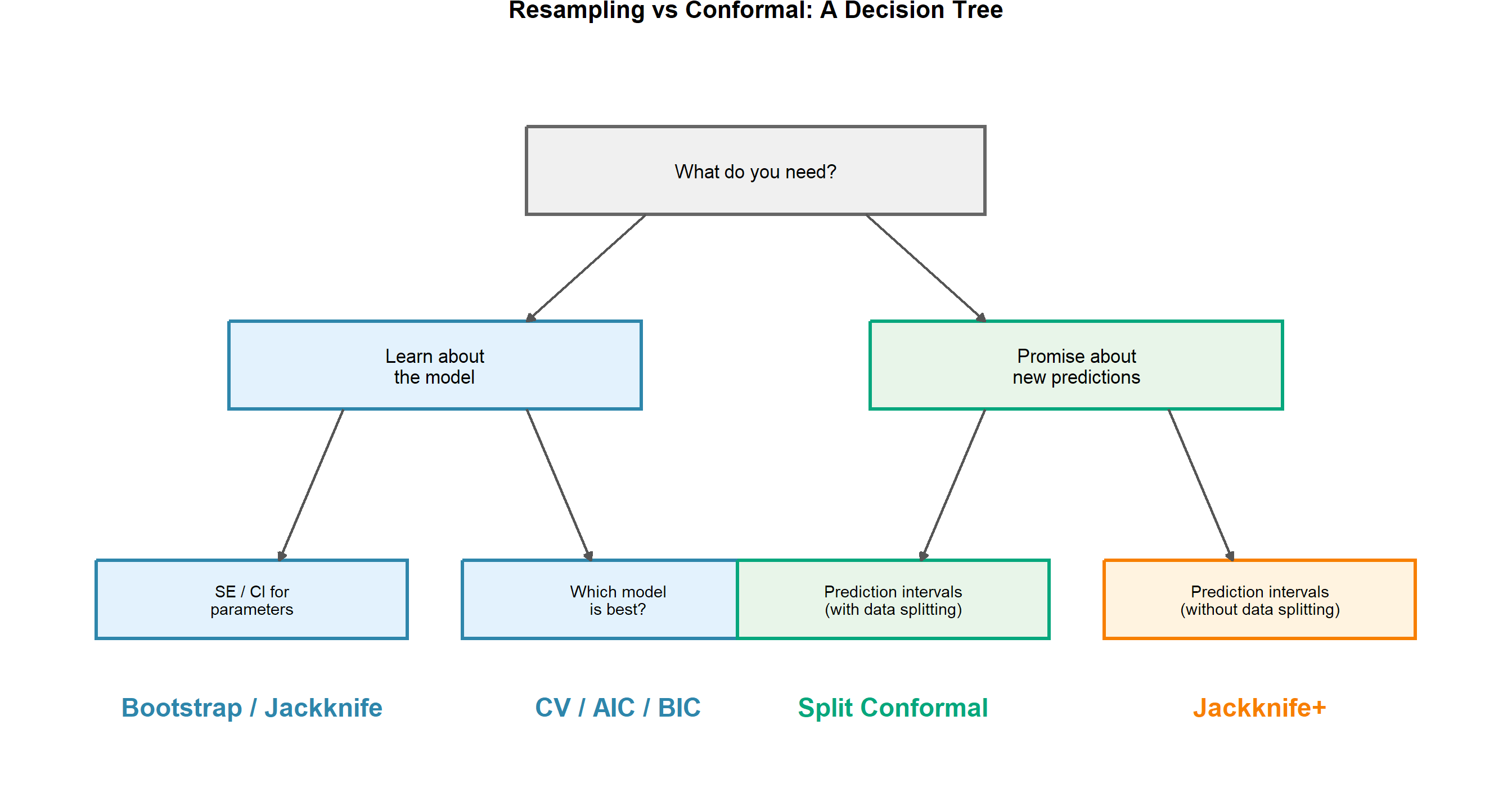
In Practice: These Paths Connect
You often use both sides sequentially. Use CV/AIC/BIC to choose your model, then wrap the chosen model in split conformal to get prediction intervals with coverage guarantees. The tree shows which tool answers which question — not that you must pick only one.
Fine Print: What the Tree Doesn’t Show
1. “Without data splitting” \(\neq\) free lunch.
Jackknife+ avoids a calibration split, but it requires \(n\) model refits (one per left-out point). For a linear model, that is cheap. For a deep network or MCMC-based model, it may be prohibitive. Split conformal requires only one refit — the price is a smaller effective training set.
2. Combining model selection with conformal requires care.
If you use CV to choose your model and the same data to calibrate conformal intervals, the calibration residuals are no longer exchangeable — they carry information about the selection step.
The clean fix: a three-way split.
\[\text{Data} \;\longrightarrow\; \underbrace{\mathcal{I}_{\text{train}}}_{\text{fit candidates}} \;+\; \underbrace{\mathcal{I}_{\text{select}}}_{\text{CV for } \lambda} \;+\; \underbrace{\mathcal{I}_{\text{calibrate}}}_{\text{conformal scores}}\]
The conformal guarantee applies to \(\mathcal{I}_{\text{calibrate}}\) because it was never touched during training or selection.
A Rule of Thumb
Any data used to make a decision about the model (which \(\lambda\), which features, which architecture) is not fresh for calibration. Keep calibration data completely untouched.
Closing: Two Sides of the Same Coin
The Week 4 Story, Complete
We asked: How do we honestly assess a model’s performance?
Information criteria (AIC, BIC) correct the optimism analytically.
Cross-validation corrects by holding out data.
Bootstrap quantifies uncertainty by resampling.
Conformal prediction guarantees coverage by exchangeability.
These are not competing schools — they are complementary tools that answer different aspects of the same question. The best practitioners know when to reach for each one.
And thanks to the paper presentations, we now know that the boundary between these worlds is porous: the jackknife+ (Barber et al., 2021) lives in both, using resampling mechanics to deliver conformal guarantees.
Looking Ahead: Classification
Weeks 4–5 recap:
- Optimism, AIC/BIC from KL divergence
- Cross-validation (LOOCV, \(k\)-fold, GCV)
- Bootstrap for uncertainty
- Conformal prediction with coverage guarantees
- Paper presentations: CQR, Jackknife+, general conformal framework
- Synthesis: resampling vs conformal paradigms
Coming up: Classification
- Bayes classifier and Bayes error rate
- Logistic regression (discriminative)
- LDA and QDA (generative)
- Decision boundaries
- Generative vs discriminative: when does each win?
Preparation
Read: ISLR Chapter 4 (Classification), ESL 4.1–4.4 (Linear Methods for Classification)