set.seed(577)
n_per <- 200
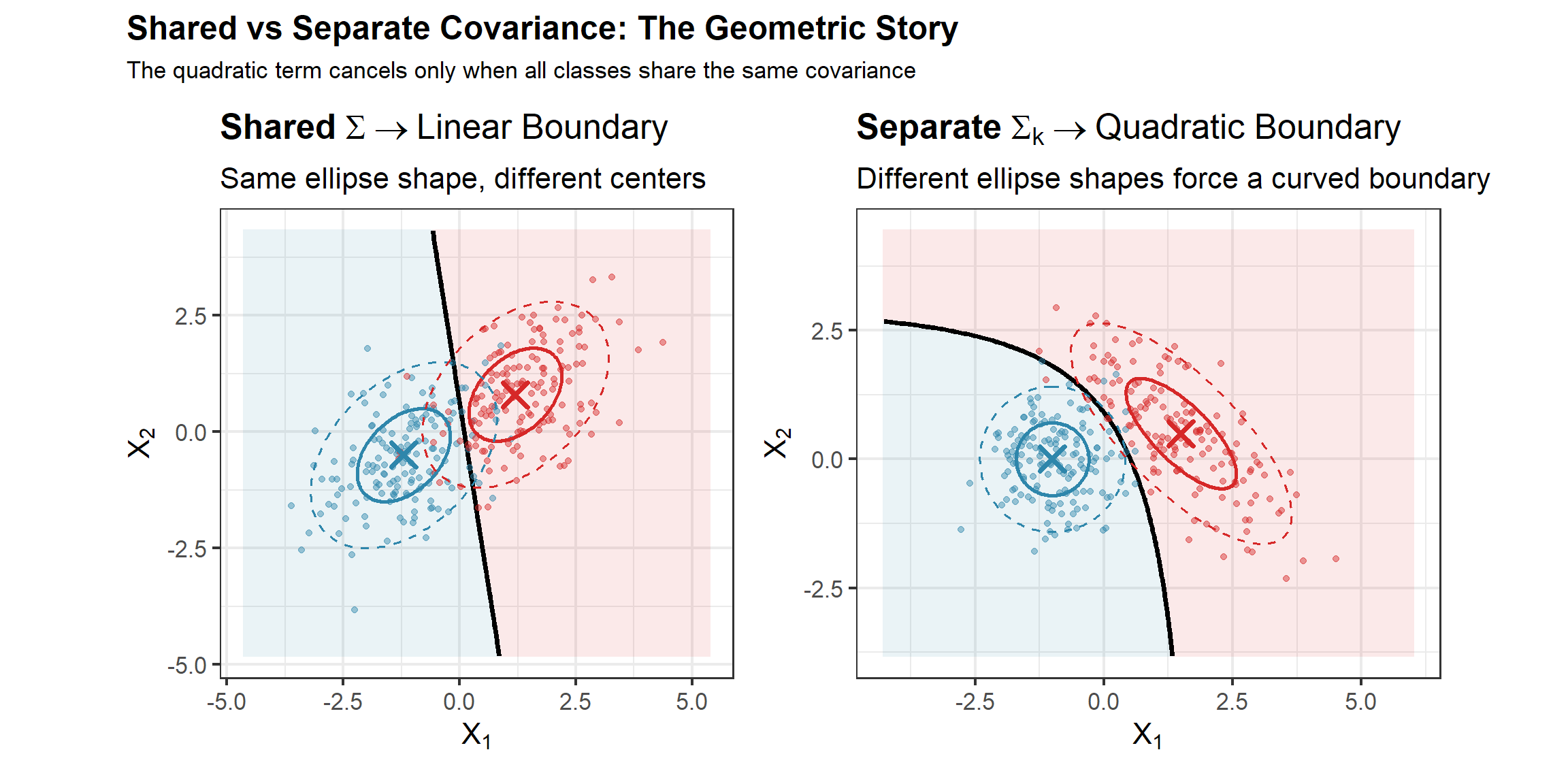
# Scenario 1: Gaussian with shared Sigma (LDA should win)
Sigma_shared <- matrix(c(1, 0.3, 0.3, 1), 2, 2)
g1_c1 <- mvrnorm(n_per, c(-1, -0.5), Sigma_shared)
g1_c2 <- mvrnorm(n_per, c(1, 0.5), Sigma_shared)
df_gauss <- tibble(x1 = c(g1_c1[,1], g1_c2[,1]),
x2 = c(g1_c1[,2], g1_c2[,2]),
class = factor(rep(0:1, each = n_per)))
# Scenario 2: Different covariances (QDA should win)
g2_c1 <- mvrnorm(n_per, c(-1, 0), matrix(c(1, 0, 0, 0.3), 2, 2))
g2_c2 <- mvrnorm(n_per, c(1, 0), matrix(c(0.3, 0, 0, 1), 2, 2))
df_diffcov <- tibble(x1 = c(g2_c1[,1], g2_c2[,1]),
x2 = c(g2_c1[,2], g2_c2[,2]),
class = factor(rep(0:1, each = n_per)))
# Scenario 3: Contaminated Gaussian (logistic should win)
# 90% clean Gaussian + 10% from 25x inflated covariance
n_clean <- round(0.9 * n_per)
n_outlier <- n_per - n_clean
contam_c1 <- rbind(
mvrnorm(n_clean, c(-1, -0.5), Sigma_shared),
mvrnorm(n_outlier, c(-1, -0.5), 25 * Sigma_shared)
)
contam_c2 <- rbind(
mvrnorm(n_clean, c(1, 0.5), Sigma_shared),
mvrnorm(n_outlier, c(1, 0.5), 25 * Sigma_shared)
)
df_contam <- tibble(x1 = c(contam_c1[,1], contam_c2[,1]),
x2 = c(contam_c1[,2], contam_c2[,2]),
class = factor(rep(0:1, each = n_per)))
# Function to compute decision boundaries for all three methods
plot_boundaries <- function(df, title_text, subtitle_text = NULL) {
grid <- expand.grid(
x1 = seq(min(df$x1) - 0.5, max(df$x1) + 0.5, length.out = 150),
x2 = seq(min(df$x2) - 0.5, max(df$x2) + 0.5, length.out = 150)
)
# Factor levels "0" < "1", so glm models P(class = "1") and
# lda posterior[,2] is also P(class = "1") --- they align correctly.
lda_fit <- lda(class ~ x1 + x2, data = df)
qda_fit <- qda(class ~ x1 + x2, data = df)
glm_fit <- glm(class ~ x1 + x2, data = df, family = binomial)
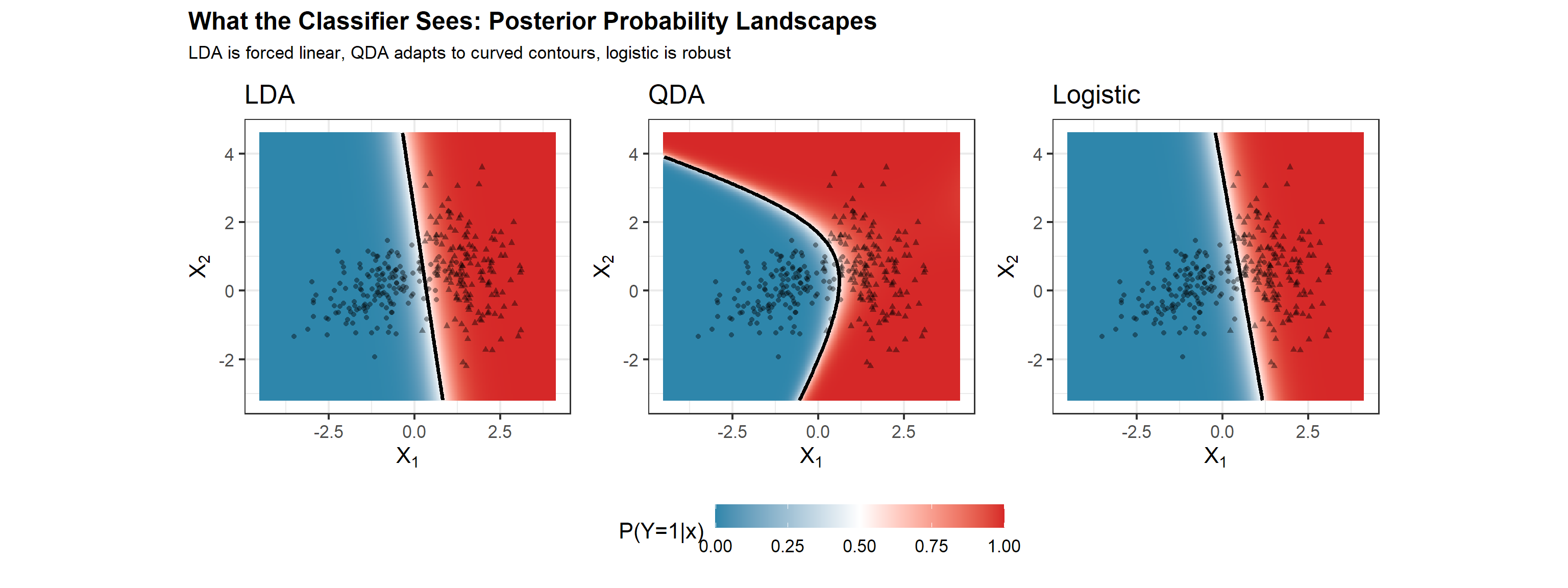
grid$lda_prob <- predict(lda_fit, grid)$posterior[,2]
grid$qda_prob <- predict(qda_fit, grid)$posterior[,2]
grid$logistic_prob <- predict(glm_fit, grid, type = "response")
ggplot(df, aes(x = x1, y = x2)) +
geom_point(aes(color = class), alpha = 0.5, size = 1.5) +
geom_contour(data = grid, aes(z = lda_prob), breaks = 0.5,
color = colors["blue"], linewidth = 1.2, linetype = "solid") +
geom_contour(data = grid, aes(z = qda_prob), breaks = 0.5,
color = colors["red"], linewidth = 1.2, linetype = "dashed") +
geom_contour(data = grid, aes(z = logistic_prob), breaks = 0.5,
color = colors["green"], linewidth = 1.2, linetype = "dotdash") +
scale_color_manual(values = c("0" = clr("blue"), "1" = clr("red")),
labels = c("Class 0", "Class 1")) +
labs(title = title_text, subtitle = subtitle_text,
x = expression(X[1]), y = expression(X[2]),
color = "True Class") +
theme_bw(base_size = 14) +
theme(legend.position = "bottom")
}
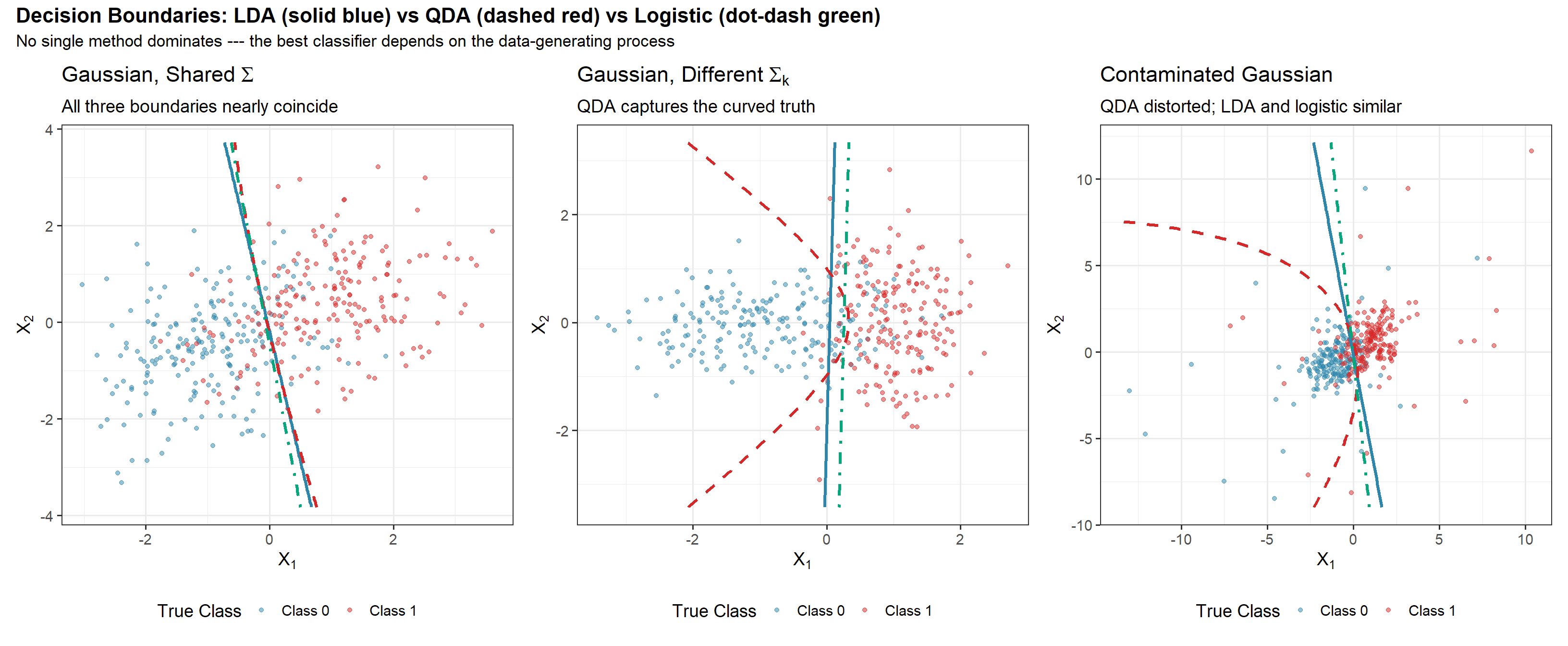
p1 <- plot_boundaries(df_gauss,
title_text = expression("Gaussian, Shared" ~ Sigma),
subtitle_text = "All three boundaries nearly coincide")
p2 <- plot_boundaries(df_diffcov,
title_text = expression("Gaussian, Different" ~ Sigma[k]),
subtitle_text = "QDA captures the curved truth")
p3 <- plot_boundaries(df_contam,
title_text = "Contaminated Gaussian",
subtitle_text = "QDA distorted; LDA and logistic similar")
p_combined <- p1 + p2 + p3 +
plot_annotation(
title = "Decision Boundaries: LDA (solid blue) vs QDA (dashed red) vs Logistic (dot-dash green)",
subtitle = "No single method dominates --- the best classifier depends on the data-generating process",
theme = theme(plot.title = element_text(size = 16, face = "bold"),
plot.subtitle = element_text(size = 13))
)
ggsave(file.path(fig_dir, "03_lda_qda_logistic_comparison.png"), p_combined,
width = 16, height = 6, dpi = 150)
p_combined